Cardiac Development and Reprogramming
While congenital heart disease (CHD) represents the most prevalent group of severe human birth defects, most of the genetic alterations that affect cardiac development and lead to CHDs remain minimally understood. Our group is interested in functional characterization of the gene regulatory landscapes driving mammalian heart development, with the goal to identify molecular mechanisms underlying cardiac disease.
Cell type-specific and/or temporally restricted expression of cardiac genes underlies tight transcriptional control by genomic cis-regulatory modules, predominantly transcriptional enhancers, which on the linear chromosome can be located at a distance (of up to 1 Mb) from target gene promoters.
We use a combination of molecular genetics, functional genomics and genome editing in embryonic stem cells and mouse embryos to study how cardiac transcription factors (TFs) with critical roles in mammalian heart development are orchestrated at the cis-regulatory level, and how disruptions in cardiac enhancer landscapes contribute to heart defects. We are also exploring the role of developmental TFs in cardiac reprogramming, with the goal to extend strategies for the regeneration of heart muscle in a disease-related context.
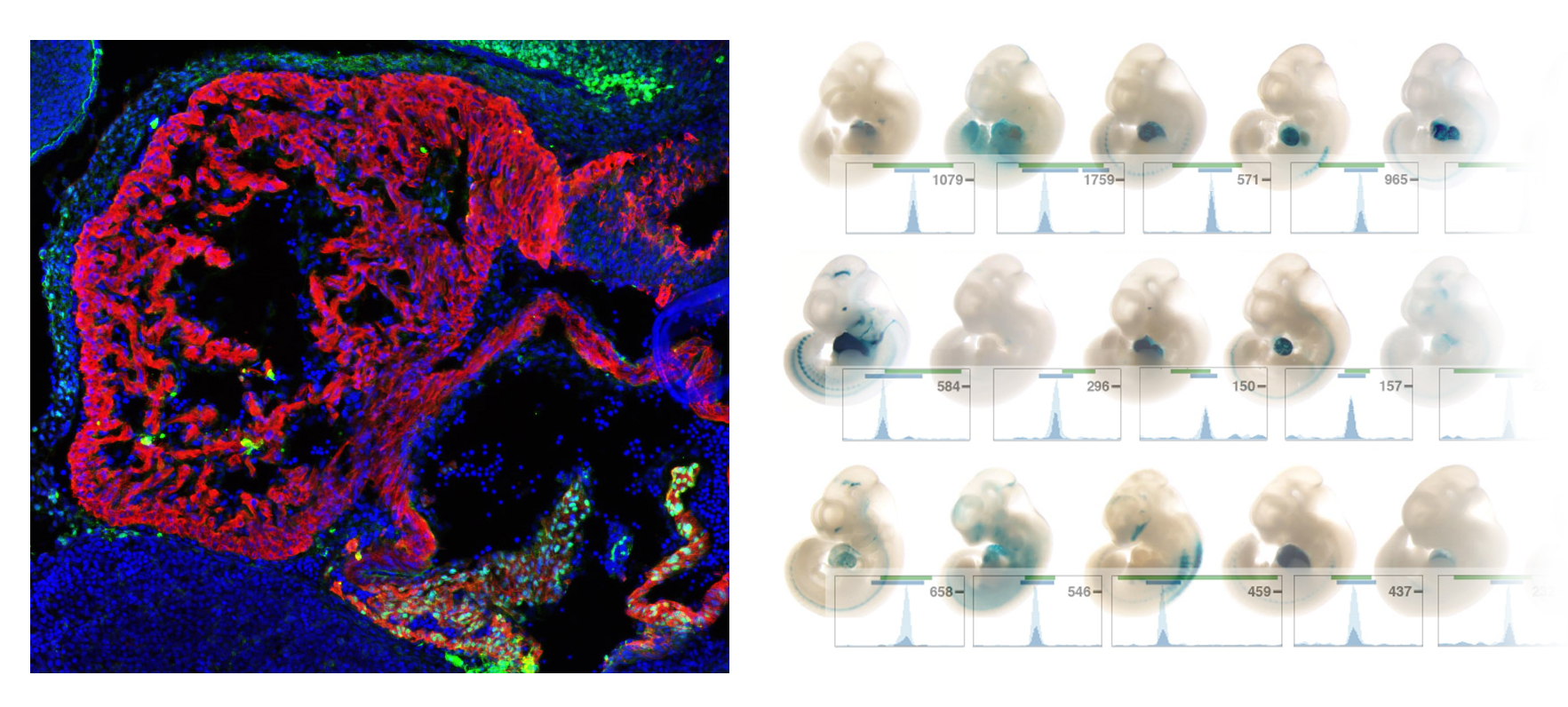
Left: Localized venous pole-restricted expression of the Shox2 transcription factor (green) essential for cardiac pacemaker function in the mouse embryonic heart myocardium (red). Right: Cardiac enhancer activities (blue) validated in transgenic LacZ reporter assays (see Vista Enhancer Browser) following genome-wide identification based on transcription factor binding and histone modifications (Laurent et al., 2017).
Contact
Marco Osterwalder, Ph.D.
SNSF Assistant Professor
University of Bern
Department for Biomedical Research (DBMR)
Murtenstrasse 24
3008 Bern
Phone: +41 (0)31 684 04 13
Email: marco.osterwalder@unibe.ch
